ReactionFlow: Visualizing Relationships between Proteins and Complexes in Biological Pathways

Background: Molecular and systems biologists are tasked with the comprehension and analysis of incredibly complex networks of biochemical interactions, called pathways, that occur within a cell. Through interviews with domain experts, we identified four common tasks that require an understanding of the causality within pathways, that is, the downstream and upstream relationships between proteins and biochemical reactions, including: visualizing downstream consequences of perturbing a protein; finding the shortest path between two proteins; detecting feedback loops within the pathway; and identifying common downstream elements from two or more proteins.
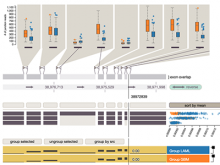
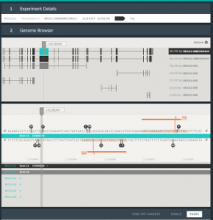
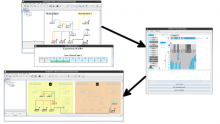
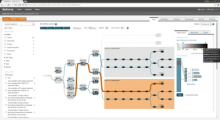
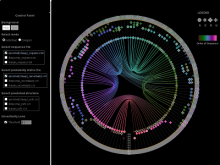
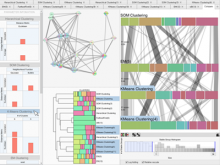
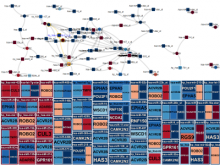
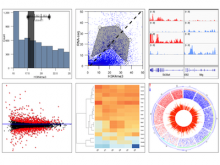
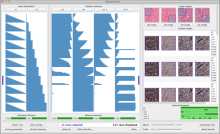
Results: We introduce ReactionFlow, a visual analytics application for pathway analysis that emphasizes the structural and causal relationships amongst proteins, complexes, and biochemical reactions within a given pathway. To support the identified causality analysis tasks, user interactions allow an analyst to filter, cluster, and select pathway components across linked views. Animation is used to highlight the flow of activity through a pathway.
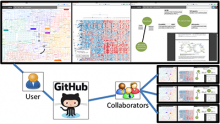
Conclusions: We evaluated ReactionFlow by providing our application to two domain experts who have significant experience with biomolecular pathways, after which we conducted a series of in-depth interviews focused on each of the four causality analysis tasks. Their feedback leads us to believe that our techniques could be useful to researchers who must be able to understand and analyze the complex nature of biological pathways. ReactionFlow is available at https://github.com/CreativeCodingLab/ReactionFlow.
BioVis 2015 Information