BioVis 2011 Paper
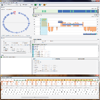
iHAT: interactive Hierarchical Aggregation Table
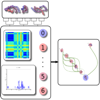
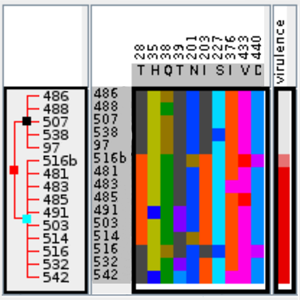
In the search for single-nucleotide polymorphisms (SNPs), genome wide association studies have become an important technique for the identification of associations between genotype and phenotype of a diverse set of sequence-based data. In this work, we present a methodology for the visual assessment of SNPs using interactive hierarchical aggregation techniques combined with methods known from traditional sequence browsers and cluster heatmaps. Our prototype tool iHAT supports the visualization of multiple sequence alignments, associated metadata, and hierarchical clusterings. Moreover, data-type dependent colormaps and aggregation strategies as well as different filtering options support the user in finding correlations between sequences and metadata. Similar to other visualizations such as parallel coordinates or heatmaps, iHAT is aimed at exploiting the human pattern-recognition ability for spotting patterns that might indicate correlation or anticorrelation. Together with its interactive features and a database backend for fast data retrieval, we consider iHAT as a prototype for a visual analytics system for genome-wide association studies.
BioVis 2011 Papers and Abstracts