Aracari: exploration of eQTL data through visualization
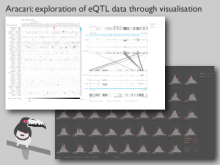
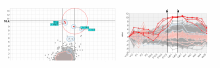
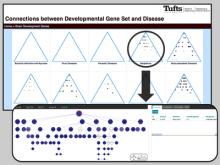
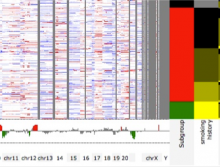
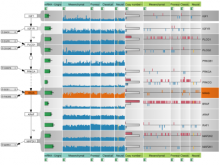
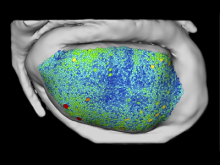
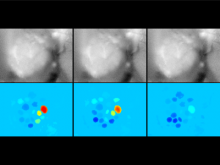
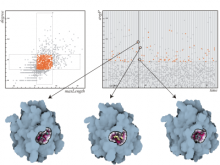
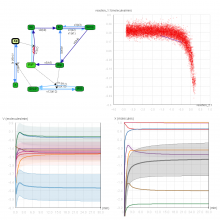
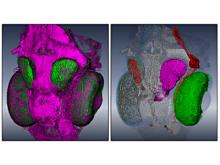
An interactive visualization tool (Aracari) was developed to analyze eQTL data. Aracari consists of two linked viewing modes: a gene expression view and a SNP view. An eQTL data set with spiked-in simulated data was analyzed; the results were assessed against the list of known spiked-in signals. Using Aracari, we were able to identify nine genes relevant to a disease while the defacto-standard biological expert’s eQTL toolkit (PLINK) could only detect six. We also identified numerous haplotype regions of relevance, highlighting a key strengths of visual analytics approach for this data.
BioVis 2012 Information