Implicit Surfaces for Interactive Graph Based Cavity Analysis of Molecular Simulations
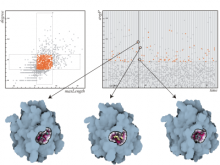
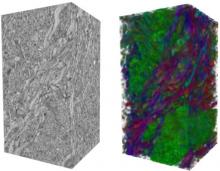
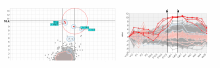
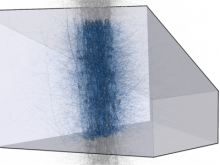
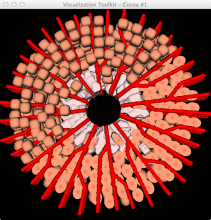
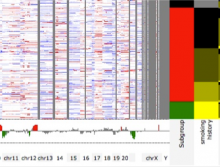
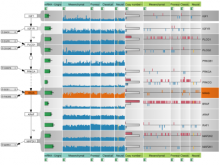
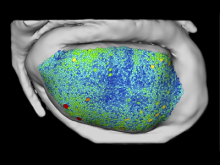
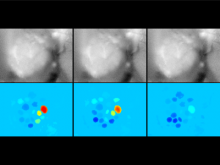
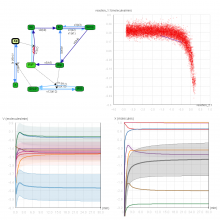
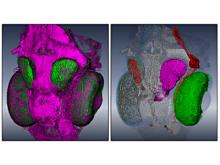
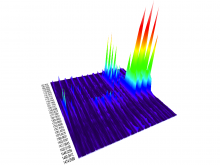
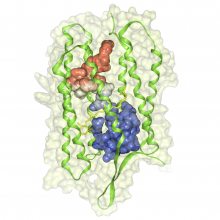
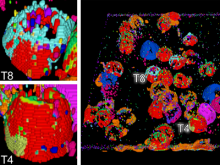
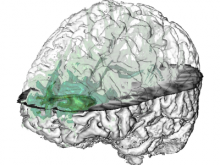
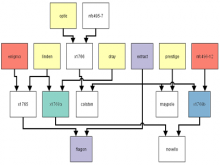
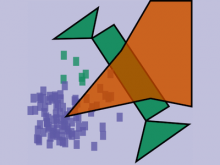
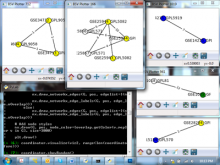
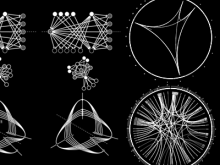
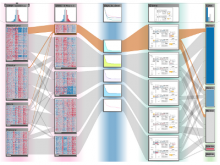
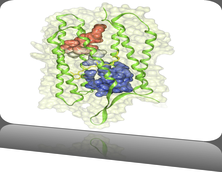
Molecular surfaces provide a suitable way to analyze and to study the evolution and interaction of molecules. \ The analysis is often concerned with visual identification of binding sites of ligands to a host macromolecule. \ We present a novel technique that is based on implicit representation, which extracts all potential binding sites and allows an advanced $3D$ visualization of these sites in the context of the molecule. \ We utilize implicit function sampling strategy to obtain potential cavity samples and graph algorithms to extract arbitrary cavity components defined by simple graphs. \ Moreover, we propose how to interactively visualize these graphs in the context of the molecular surface. \ We also introduce a system of linked views depicting various graph parameters that are used to perform a more elaborative study on created graphs.
BioVis 2012 Information