MedSavant: Visual Analytics for Genetic Variation Datasets
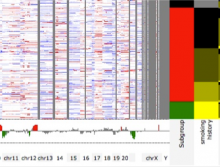
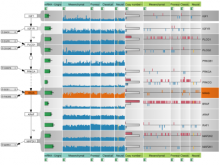
High Throughput Sequencing (HTS) technologies have revolutionized the speed and economy with which genomic information can be obtained, and are providing a means for deep cataloguing of human variation. One of the most challenging problems is in identifying those few genetic variants (among potentially millions that are predicted per sequenced individual) that are actually causal in genetic disease. For this purpose we introduce MedSavant: a software platform for accelerating the identification of disease-causing genetic variants found in population sequencing studies by enabling complex and dynamic querying of patient data, and by delivering informative visualizations of the results.
BioVis 2012 Information