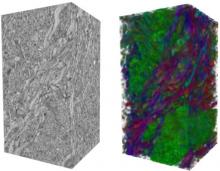
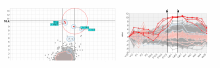
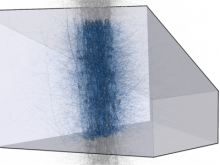
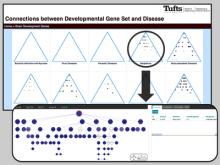
Visualizing Cells and their Connectivity Graphs for CompuCell3D
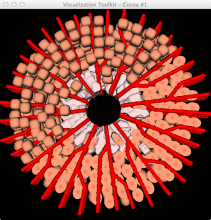

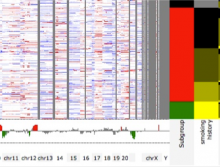
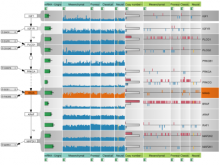
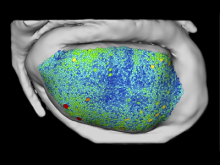
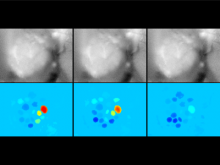
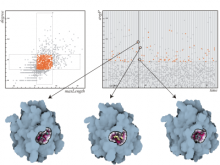
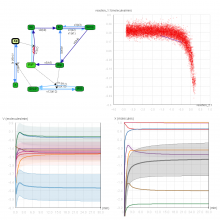
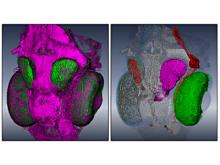
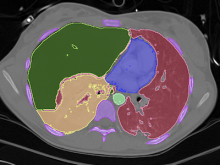
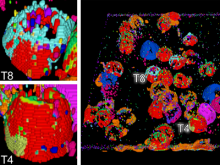
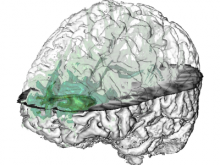
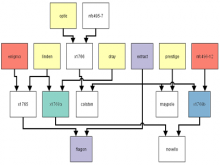
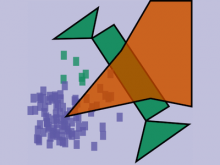
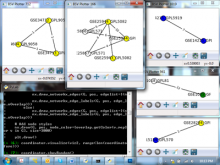
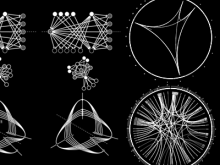
Developing models that simulate the behavior of different types of interacting biological cells can be a very time consuming and error prone task. CompuCell3D is an open source application that addresses this challenge. It provides interactive and customizable visualizations that help a user detect when a model is producing the desired behavior and when it is failing. It also allows for high quality image generation for publications and presentations. CompuCell3D uses the Python programming language which allows for easy extensions. Examples are provided for performing graph analyses of cell connectivity.
BioVis 2012 Information